For quotations, please use our online quotation form, and you may also contact us by
sales@neoscientific.com
+1-888.733.6849
+1-617.299.7367 (Int’l)
+1-888.733.6849
+1-617.299.7367 (Int’l)
| Reactivity | Human Mouse Rat |
| Tested applications | WB IHC IF |
| Recommended Dilution | WB 1:500 - 1:2000 IHC 1:50 - 1:200 IF 1:50 - 1:200 |
| Calculated MW | 166kDa |
| Observed MW | Refer to Figures |
| Immunogen | Recombinant protein of human FANCD2 |
| Storage Buffer | Store at -20℃. Avoid freeze / thaw cycles. Buffer: PBS with 0.02% sodium azide, 50% glycerol, pH7.3. |
| Concentration | e |
| Synonym | DKFZp762A223; FA-D2; FA4; FACD; FAD; FAD2; FANCD; FLJ23826; |
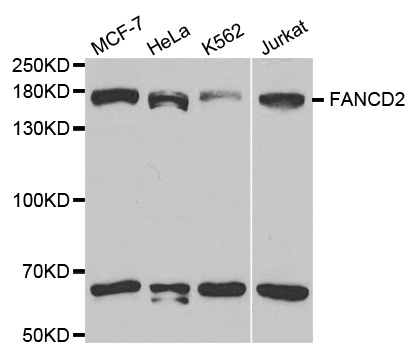
Western blot analysis of extracts of various cell lines, using FANCD2 antibody.
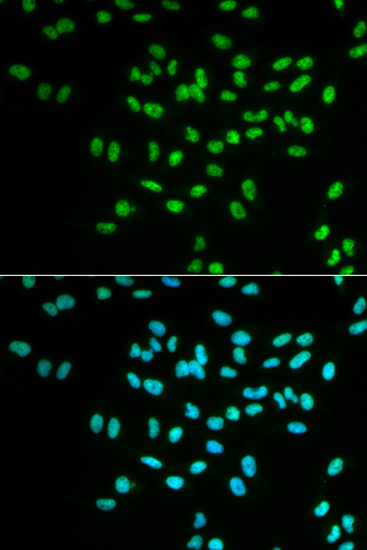
Immunofluorescence analysis of MCF7 cell using FANCD2 antibody. Blue: DAPI for nuclear staining.
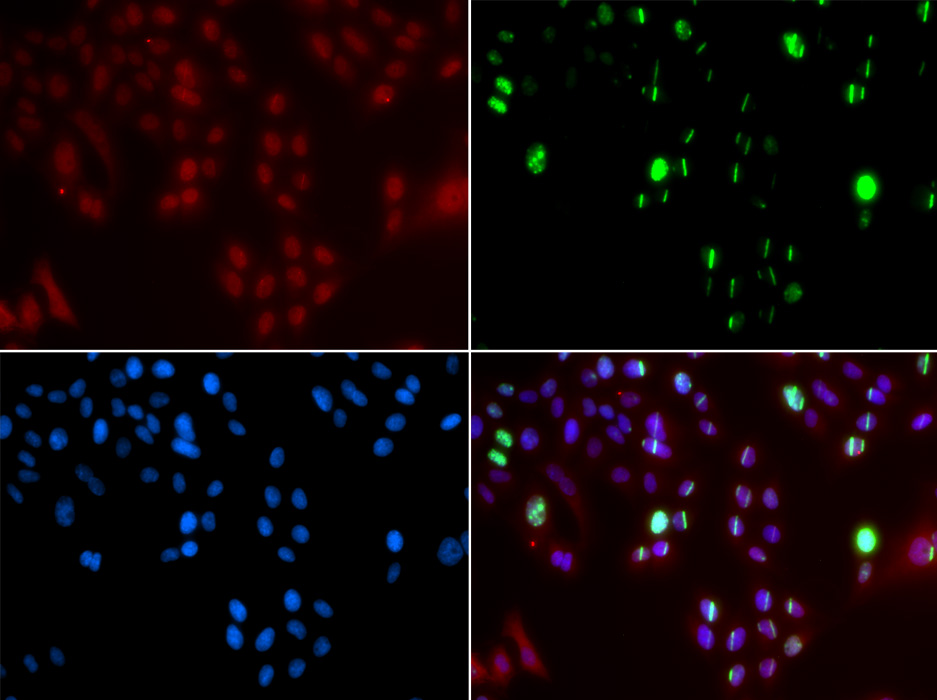
Immunofluorescence analysis of GFP-RNF168 transgenic U2OS cell using FANCD2 antibody. Green:GFP-RNF168 fusion protein expression for DNA damage marker.Blue: DAPI for nuclear staining.RNF168(GFP) can be used to mark cells damaged by UV-A laser for they always gather around DNA damage region.
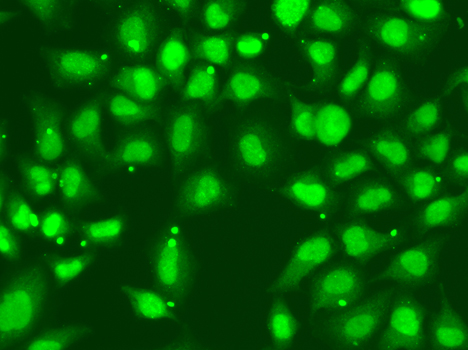
Immunofluorescence analysis of A549 cell using FANCD2 antibody.
The Fanconi anemia complementation group (FANC) currently includes FANCA, FANCB, FANCC, FANCD1 (also called BRCA2), FANCD2, FANCE, FANCF, FANCG, FANCI, FANCJ (also called BRIP1), FANCL, FANCM and FANCN (also called PALB2). The previously defined group FANCH is the same as FANCA. Fanconi anemia is a genetically heterogeneous recessive disorder characterized by cytogenetic instability, hypersensitivity to DNA crosslinking agents, increased chromosomal breakage, and defective DNA repair. The members of the Fanconi anemia complementation group do not share sequence similarity; they are related by their assembly into a common nuclear protein complex. This gene encodes the protein for complementation group D2. This protein is monoubiquinated in response to DNA damage, resulting in its localization to nuclear foci with other proteins (BRCA1 AND BRCA2) involved in homology-directed DNA repair. Alternative splicing results in two transcript variants encoding different isoforms. [provided by RefSeq, Jul 2008]
N/A